图像处理实践 |
您所在的位置:网站首页 › 色彩识别测试方法有哪些 › 图像处理实践 |
图像处理实践
|
水果图像的识别与分类
1 数据获取与数据集介绍
数据来源:公开水果数据集fruit-360,包含几十种水果的彩色图片,图片格式为100*100像素,训练集中,每种水果都有上百张各种角度拍摄的照片。可以通过对图像的预处理、特征提取,并构建分类器对于水果照片进行分类。 数据集可从Github上下载:https://github.com/Horea94/Fruit-Images-Dataset 2 预处理与特征提取 2.1 相关库的导入对于图像分类问题,我们使用python-opencv的相关模块来对于图像进行预处理,同时,使用scikit-image来提取图像中的一些特征,这里主要尝试使用HOG (Histogram of Oriented Gridients) 来描述图像中局部纹理特征,分类器我们使用sklearn中的SVM、kNN、决策树等模型。 import numpy as np import cv2 import glob import os import matplotlib.pyplot as plt import string from mlxtend.plotting import plot_decision_regions from mpl_toolkits.mplot3d import Axes3D from sklearn.decomposition import PCA from sklearn.preprocessing import StandardScaler from sklearn.neighbors import KNeighborsClassifier from sklearn.tree import DecisionTreeClassifier from sklearn.model_selection import train_test_split, cross_val_score from sklearn.utils.multiclass import unique_labels from sklearn import metrics from sklearn.svm import SVC # HOG import json from skimage import color from skimage.feature import hog from sklearn import svm from sklearn.metrics import classification_report,accuracy_score from subprocess import check_output dim = 100 2.2 数据集导入数据集包括多个文件夹,每个文件夹中是同一种水果,这里我们定义一个函数方便我们进行数据导入。 def GetFruit(fruitlist,datatype,print_n=False,k_fold=False): images=[] labels=[] val=['train','test'] if not k_fold: PATH="E:/kaggle/fruit-360/"+datatype+"/" for i,fruit in enumerate(fruitlist): p=PATH+fruit j=0 for image_path in glob.glob(os.path.join(p,"*.jpg")): image=cv2.imread(image_path,cv2.IMREAD_COLOR) image=cv2.resize(image,(dim,dim)) image=cv2.cvtColor(image,cv2.COLOR_RGB2BGR) images.append(image) labels.append(i) j+=1 if(print_n): print("There are",j,"",datatype.upper()," images of ",fruitlist[i].upper()) images=np.array(images) labels=np.array(labels) return images,labels else: for v in val: PATH="E:/kaggle/fruit-360/"+v+"/" for i,fruit in enumerate(fruitlist): p=PATH+fruit j=0 for image_path in glob.glob(os.path.join(p,"*.jpg")): image=cv2.imread(image_path,cv2.IMREAD_COLOR) image=cv2.resize(image,(dim,dim)) image=cv2.cvtColor(image,cv2.COLOR_RGB2BGR) images.append(image) labels.append(i) j+=1 images=np.array(images) labels=np.array(labels) return images,labels def GetAll(): fruitlist=[] for fruit_path in glob.glob("E:/kaggle/fruit-360/train/*"): fruit=frui_path.split("/")[-1] fruitlist.append(fruit) return fruits同时,我们还希望在图像识别的过程中明白算法中间发生了什么事情,因此可以通过一定的可视化操作来实现这一点,这里定义相关的可视化函数: def getClassNumber(y): v =[] i=0 count = 0 for index in y: if(index == i): count +=1 else: v.append(count) count = 1 i +=1 v.append(count) return v def plotPrincipalComponents(X, dim): v = getClassNumber(y_train) colors = 'orange', 'purple', 'r', 'c', 'm', 'y', 'k', 'grey', 'b', 'g' markers = ['o', 'x' , 'v', 'd'] tot = len(X) start = 0 if(dim == 2): for i,index in enumerate(v): end = start + index plt.scatter(X[start:end,0],X[start:end,1] , color=colors[i%len(colors)], marker=markers[i%len(markers)], label = fruitlist[i]) start = end plt.xlabel('PC1') plt.ylabel('PC2') if(dim == 3): fig = plt.figure() ax = fig.add_subplot(111, projection='3d') for i,index in enumerate(v): end = start + index ax.scatter(X[start:end,0], X[start:end,1], X[start:end,2], color=colors[i%len(colors)], marker=markers[i%len(markers)], label = fruitlist[i]) start = end ax.set_xlabel('PC1') ax.set_ylabel('PC2') ax.set_zlabel('PC3') plt.legend(loc='lower left') plt.xticks() plt.yticks() plt.show() # 绘制混淆矩阵 def plot_confusion_matrix(y_true, y_pred, classes, normalize=False, title=None, cmap=plt.cm.Blues): """ This function prints and plots the confusion matrix. Normalization can be applied by setting `normalize=True`. """ if not title: if normalize: title = 'Normalized confusion matrix' else: title = 'Confusion matrix, without normalization' # Compute confusion matrix cm = metrics.confusion_matrix(y_true, y_pred) # Only use the labels that appear in the data classes = unique_labels(y_true, y_pred) if normalize: cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis] fig, ax = plt.subplots() im = ax.imshow(cm, interpolation='nearest', cmap=cmap) ax.figure.colorbar(im, ax=ax) # We want to show all ticks... ax.set(xticks=np.arange(cm.shape[1]), yticks=np.arange(cm.shape[0]), # ... and label them with the respective list entries xticklabels=fruitlist, yticklabels=fruitlist, title=title, ylabel='True label', xlabel='Predicted label') # Rotate the tick labels and set their alignment. plt.setp(ax.get_xticklabels(), rotation=45, ha="right", rotation_mode="anchor") # Loop over data dimensions and create text annotations. fmt = '.2f' if normalize else 'd' thresh = cm.max() / 2. for i in range(cm.shape[0]): for j in range(cm.shape[1]): ax.text(j, i, format(cm[i, j], fmt), ha="center", va="center", color="white" if cm[i, j] > thresh else "black") fig.tight_layout() return cm,ax 2.3 数据查看与编码取一张图片,查看其RGB直方图,这一点可以直接由cv模块中的相关函数来完成。从直方图上我们能够看到图像数据的频率分布,便于我们检测图像的明度和色彩的变化趋势。 sample = cv2.imread("E:/kaggle/fruit-360/train/Kiwi/0_100.jpg",cv2.IMREAD_COLOR) plt.hist(sample.ravel(), bins=256, range=[0, 256]); plt.show() color = ('b', 'g', 'r') for i, col in enumerate(color): histr = cv2.calcHist([sample], [i], None, [256], [0, 256]) plt.plot(histr, color=col) plt.xlim([0, 256]) plt.show()
可以看到,图像的曝光程度正常,直方图没有明显偏左或者偏右。同时GRB三条曲线的偏离不是很大,说明白平衡没有较大问题。 为了简单起见,考虑二分类问题,这里我们需要区分菠萝和猕猴桃(选择这两种水果是因为从外表来看颜色比较相似,想要区分不会那么容易) # Binary classification fruitlist=['Pineapple','kiwi'] # Get image and Labels X_train_raw,y_train=GetFruit(fruitlist,'train',print_n=True,k_fold=False) X_test_raw,y_test=GetFruit(fruitlist,'test',print_n=True,k_fold=False) # Get data for k-fold X_raw,y=GetFruit(fruitlist,'',print_n=True,k_fold=True)将所有图片储存在列表里,接着提取图片的hog特征。HOG先计算图片某一区域中不同方向上梯度的值,然后进行累积,得到直方图,这个特征能够直接输入到分类器当中实现图像分类。 from skimage import feature ppc=16 hog_features_train = [] hog_features_test = [] hog_images_train= [] hog_images_test=[] for image in X_train_raw: fd,hog_image = feature.hog(image, orientations=8, pixels_per_cell=(ppc,ppc),cells_per_block=(4, 4),block_norm= 'L2',visualize=True) hog_images_train.append(hog_image) hog_features_train.append(fd) for image in X_test_raw: fd,hog_image = feature.hog(image, orientations=8, pixels_per_cell=(ppc,ppc),cells_per_block=(4, 4),block_norm= 'L2',visualize=True) hog_images_test.append(hog_image) hog_features_test.append(fd) # Scale data Images scaler=StandardScaler() X_train=scaler.fit_transform([i.flatten() for i in X_train_raw]) X_test=scaler.fit_transform([i.flatten() for i in X_test_raw]) X_train_hog=scaler.fit_transform([i.flatten() for i in hog_images_train]) X_test_hog=scaler.fit_transform([i.flatten() for i in hog_images_test]) X=scaler.fit_transform([i.flatten() for i in X_raw]) There are 490 TRAIN images of PINEAPPLE There are 466 TRAIN images of KIWI There are 166 TEST images of PINEAPPLE There are 156 TEST images of KIWI可以看到,菠萝的训练集和测试集分别有490、166张图片,猕猴桃的训练集分别有466、156张图片。 print("Shape of the data:") print((X_train.shape,y_train.shape)) print((X_test.shape,y_test.shape)) print(X_train_hog.shape) print(X_test_hog.shape) print("\nData sample:") print((X_train[0],y_train[0])) print((X_train_hog[0],y_train[0])) # x中是数据,每张图片由100*100像素,3条RGB通道构成,一共是30000维的向量构成 # y中是预测结果,二分类中被编码为0和1 Shape of the data: ((956, 30000), (956,)) ((322, 30000), (322,)) (956, 10000) (322, 10000) Data sample: (array([-1.35474176, 0.24001185, 0.26975539, ..., 0. , 0. , 0. ]), 0) (array([0., 0., 0., ..., 0., 0., 0.]), 0)编码后的图像每张图片被转化为一个30000维的向量,包括100*100像素点的三个RGB通道,同时我们将0-255的值进行尺度变化,转化为-1至1的浮点数。 # 查看数据样例 def plot_image_grid(images, nb_rows, nb_cols, figsize=(15, 15)): assert len(images) == nb_rows*nb_cols, "Number of images should be the same as (nb_rows*nb_cols)" fig, axs = plt.subplots(nb_rows, nb_cols, figsize=figsize) n = 0 for i in range(0, nb_rows): for j in range(0, nb_cols): axs[i, j].axis('off') axs[i, j].imshow(images[n]) n += 1 print(fruitlist) plot_image_grid(X_train_raw[0:100], 10, 10) plot_image_grid(X_train_raw[490:590], 10, 10) ['Pineapple', 'kiwi']
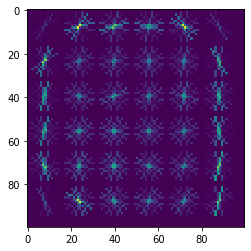
可以看到,数据集中的水果图像有各个角度拍摄的,同时我们也可以把单张图片的原图和提取HOG特征后的图像绘制出来: plt.imshow(X_train_raw[1])
提取HOG特征之后,图像在关键的点位中只剩下了特征向量的部分,这里可以看到图像受到光照、角度、颜色的影响明显变小,这样的特征提取有利于我们对于图像进行分类。 3 构建基本的SVM分类器使用sklearn模块中的SVM分类器,这里我们将提取特征后的图像X_train_hog输入到分类器当中进行训练,同时基于测试集X_test_hog进行模型效果的测试。 # linear SVM using hog features svm=SVC(gamma='auto',kernel='linear',probability=True) svm.fit(X_train_hog,y_train) y_pred=svm.predict(X_test_hog) # Evaluation precision=metrics.accuracy_score(y_pred,y_test)*100 print("Accuracy with SVM: {0:.2f}%".format(precision)) cm,_=plot_confusion_matrix(y_test,y_pred,classes=y_train,normalize=True,title="Normalized confusion matrix") plt.show() # calculate FPR and TPR probs=svm.predict_proba(X_test_hog) probs=probs[:,1] svm_fpr,svm_tpr,thresholds=metrics.roc_curve(y_test,probs) svm_auc=metrics.roc_auc_score(y_test,probs) Accuracy with SVM: 100.00%
可以看到,将图像提取出的特征用SVM进行分类之后准确率达到了100%,同时我们可以看到在测试集上的混淆矩阵,这说明用这种方法进行分类是相当有效的。 4 更多分类模型的尝试 4.1 使用PCA进行降维PCA可以通过线性变换,重整高维数据,提取其中的重要部分,忽略其中无关紧要的部分,这里我们使用PCA对于图片进行处理,同时对中间过程进行可视化。 # 使用PCA提取特征 pca = PCA(n_components=3) dataIn3D = pca.fit_transform(X_train) plotPrincipalComponents(dataIn3D, 3)
可以看到,两种水果的前三个主成分在空间中的分布有所不同,因此我们可以根据数据点在空间位置的不同来构建分类器。 # 展示PCA内部细节的模块 def showPCA(image,X2, X10, X50): fig = plt.figure(figsize=(15,15)) ax1 = fig.add_subplot(1,4,1) ax1.axis('off') ax1.set_title('Original image') plt.imshow(image) ax1 = fig.add_subplot(1,4,2) ax1.axis('off') ax1.set_title('50 PC') plt.imshow(X50) ax1 = fig.add_subplot(1,4,3) ax1.axis('off') ax1.set_title('10 PC') plt.imshow(X10) ax2 = fig.add_subplot(1,4,4) ax2.axis('off') ax2.set_title('2 PC') plt.imshow(X2) plt.show() def computePCA(n, im_scaled, image_id): pca = PCA(n) principalComponents = pca.fit_transform(im_scaled) im_reduced = pca.inverse_transform(principalComponents) newImage = scaler.inverse_transform(im_reduced[image_id]) return newImage def showVariance(X_train): #Compute manually the principal components cov_matr=np.dot(X_train, X_train.T) eigval,eigvect=np.linalg.eig(cov_matr) index=np.argsort(eigval)[::-1] #take in order the index of ordered vector (ascending order) #eigvect[:,i] is associated to eigval[i] so eigvect=eigvect[:,index] eigval=eigval[index] n_PC=[] var_explained=[] var_temp=[] var_tmp=0 for i in range(10): var_tmp=var_tmp+eigval[i] n_PC.append(i) var_temp.append(eigval[i]/(eigval.sum())*100) var_explained.append(var_tmp/(eigval.sum())*100) fig, ax = plt.subplots(figsize=(8,8)) ind = np.arange(10) width = 0.35 # the width of the bars p1 = ax.bar(ind, var_temp, width, color='orange') p2 = ax.bar(ind + width, var_explained, width, color='blue') ax.legend((p1[0], p2[0]), ('Individual explained variance', 'Cumulative explained variance')) ax.set_title('Variance explained using PCs') ax.set_xticks(ind + width / 2) ax.set_xticklabels(('1', '2', '3', '4', '5', '6', '7', '8', '9', '10')) plt.xlabel('Number of PC') plt.ylabel('Variance exaplained in %') ax.autoscale_view() plt.show() image_id = 2 image = X_train_raw[image_id] #Compute PCA X_2 = computePCA(2, X_train,image_id) X_10 = computePCA(10, X_train,image_id) X_50 = computePCA(50, X_train,image_id) #Reshape in order to plot images X2 = np.reshape(X_2, (dim,dim,3)).astype(int) X10 = np.reshape(X_10, (dim,dim,3)).astype(int) X50 = np.reshape(X_50, (dim,dim,3)).astype(int) #Plot showPCA(image, X2, X10, X50) Clipping input data to the valid range for imshow with RGB data ([0..1] for floats or [0..255] for integers). Clipping input data to the valid range for imshow with RGB data ([0..1] for floats or [0..255] for integers). Clipping input data to the valid range for imshow with RGB data ([0..1] for floats or [0..255] for integers).
可以看到,选取的主成分越少,图像就变得越为抽象,选取的主成分越多,图像就越接近原始的图像,展现的细节也越多。 showVariance(X_train)
可以看到,PCA降维后再次输入SVM也能保持几乎100%的分类正确率。我们可以画出只选取前两个主成分的PCA降维分类的图像: pca = PCA(n_components=2) X_train2D = pca.fit_transform(X_train) X_test2D = pca.fit_transform(X_test) svm.fit(X_train2D, y_train) test_predictions = svm.predict(X_test2D) precision = metrics.accuracy_score(test_predictions, y_test) * 100 print("Accuracy with SVM considering only first 2PC: {0:.2f}%".format(precision)) #Plotting decision boundaries plot_decision_regions(X_train2D, y_train, clf=svm, legend=1) plt.xlabel('PC1') plt.ylabel('PC2') plt.title('Linear SVM Decision Boundaries') plt.show() Accuracy with SVM considering only first 2PC: 98.14%
另外也可以尝试使用核PCA: svm_with_kernel = SVC(gamma=0.01, kernel='rbf', probability=True) svm_with_kernel.fit(X_train2D, y_train) y_pred = svm_with_kernel.predict(X_test2D) precision = metrics.accuracy_score(y_pred, y_test) * 100 print("Accuracy with Not-Linear SVM considering only first 2PC: {0:.2f}%".format(precision)) #Plotting decision boundaries plot_decision_regions(X_train2D, y_train, clf=svm_with_kernel, legend=1) plt.xlabel('PC1') plt.ylabel('PC2') plt.title('Kernel SVM Decision Boundaries') plt.show() Accuracy with Not-Linear SVM considering only first 2PC: 64.91%
可以看到使用rbf核的PCA在测试集的准确率不及线性核,从图像中也可以看出,实现分类的并不是一条直线,而是不规则的边界。 4.3 使用kNN分类器 # knn knn = KNeighborsClassifier(n_neighbors=2) knn.fit(X_train, y_train) y_pred = knn.predict(X_test) #Evaluation precision = metrics.accuracy_score(y_pred, y_test) * 100 print("Accuracy with K-NN: {0:.2f}%".format(precision)) cm , _ = plot_confusion_matrix(y_test, y_pred, classes=y_train, normalize=True, title='Normalized confusion matrix') plt.show() # calculate the FPR and TPR for all thresholds of the classification probs = knn.predict_proba(X_test) probs = probs[:, 1] knn_fpr, knn_tpr, thresholds = metrics.roc_curve(y_test, probs) knn_auc = metrics.roc_auc_score(y_test, probs) Accuracy with K-NN: 99.69%
可以看到,kNN分类器能够达到99%的精度,同理,也可以将决策边界给画出来从而理解数据本身: #K-NN + PCA knn.fit(X_train2D, y_train) y_pred = knn.predict(X_test2D) precision = metrics.accuracy_score(y_pred, y_test) * 100 print("Accuracy with K-NN considering only first 2PC: {0:.2f}%".format(precision)) #Plotting decision boundaries plot_decision_regions(X_train2D, y_train, clf=knn, legend=1) plt.xlabel('PC1') plt.ylabel('PC2') plt.title('K-NN Decision Boundaries') plt.show() Accuracy with K-NN considering only first 2PC: 98.14%
可以看到,使用决策树分类器能够达到95%精确率,我们也能将决策边界绘制出来: #DECISION TREE + PCA tree = tree.fit(X_train2D,y_train) y_pred = tree.predict(X_test2D) precision = metrics.accuracy_score(y_pred, y_test) * 100 print("Accuracy with Decision Tree considering only first 2PC: {0:.2f}%".format(precision)) #Plotting decision boundaries plot_decision_regions(X_train2D, y_train, clf=tree, legend=1) plt.xlabel('PC1') plt.ylabel('PC2') plt.title('Decision Tree Decision Boundaries') plt.show() Accuracy with Decision Tree considering only first 2PC: 97.52%
参考文章(kernel): Training svm classifier with hog features. (Manik Galkissa) Fruit Classification: PCA, SVM, KNN, Decision Tree. (Walter Maffione) |
【本文地址】
今日新闻 |
推荐新闻 |