净重新分类指数(NRI)的计算方法 |
您所在的位置:网站首页 › 净重的公式 › 净重新分类指数(NRI)的计算方法 |
净重新分类指数(NRI)的计算方法
 1. 背景知识 净重新分类指数(NRI)这个指标最初用于评价诊断试验中新的诊断指标较旧诊断指标把研究对象进行正确分类在数量上的变化。既然可以评估诊断试验预测的准确度,那自然也可以用于判断预测模型的准确度,从已发表的临床研究文献来看,NRI这个指标更广泛应用于比较两个预测模型的准确度。前文已经述及比较两个预测模型的准确性或者区分度的方法,一般计算C-statistics,或者叫ROC曲线下面积AUC,而C-statistics或AUC具有一定的局限性: (1)C-statistics/AUC不够敏感,当我们想要在原有模型中引入新的指标后,观察模型的预测能力是否有所提高,此时新加入的指标有时很难显著改善C-statistics/AUC,其增量往往并不明显; (2)C-statistics/AUC的专业意义不容易理解,很难转化为恰当的临床解释。而NRI很好的克服了这两个不足。 2. 计算原理 我们首先以二分类诊断指标为例讲解NRI大致计算原理,然后再将其扩展应用于预测模型预测能力的量化比较,前者一般手动计算即可,后者计算需借助统计软件。简单来讲,旧诊断指标会把研究对象分类为患者和非患者,而新的诊断指标会把研究对象再重新分类为患者和非患者。此时比较新、旧诊断指标对于研究人群的分类变化,就会发现有一部分研究对象,原本在旧诊断指标中被错分,但在新诊断指标中得到了正确划分;同样也有一部分研究对象,原本在旧诊断指标中分类正确,但在新诊断指标中却被错分,因此研究对象的分类在新、旧诊断指标中会发生变化,我们利用这种重新分类的变化,来计算净重新分类指数NRI[1,2]。文字读起来确实有点绕,可参考下面计算过程帮助消化。 首先我们将研究对象按照金标准的诊断结果分为患病和未患病两组,然后分别在这两个分组下,根据新、旧诊断指标的预测分类结果,整理成两个配对的四格表,如下表1.和表2.所示。 表1. 患病租重新分类表 患病组(N1)新指标旧指标阳性阴性阳性a1b1阴性c1d1表2. 无病组重新分类表 无病组(N2)新指标旧指标阳性阴性阳性a2b2阴性c2d2我们主要关注被重新分类的研究对象,从表中可以看出,在患病组(总数为N1),新诊断指标分类正确而旧指标分类错误的有c1个人,新指标分类错误而旧指标分类正确的有b1个人,那么新模型相对于旧模型来说,正确分类提高的比例为(c1-b1)/N1。同理,在非患者组(总数为N2),新诊断指标分类正确而旧指标分类错误的有b2个人,新指标分类错误而旧指标分类正确的有c2个人,那么新诊断指标相对于旧指标正确分类提高的比例为(b2-c2)/ N2。最后,综合患者组和非患者组的结果,新模型与旧模型相比,净重新分类指数NRI= (c1-b1)/N1+(b2-c2)/N2,一般称作绝对NRI。 若NRI>0,则为正改善,说明新指标比旧指标的预测能力有所改善;若NRI= 0.4 0 2 84 ## Reclassification Table for case: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 7 0 0 ## < 0.4 0 8 0 ## >= 0.4 0 2 71 ## Reclassification Table for control: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 103 3 0 ## < 0.4 3 22 0 ## >= 0.4 0 0 13 ## NRI estimation: ## Point estimates: ## Estimate ## NRI -0.02272727 ## NRI+ -0.02272727 ## NRI- 0.00000000 ## Pr(Up|Case) 0.00000000 ## Pr(Down|Case) 0.02272727 ## Pr(Down|Ctrl) 0.02083333 ## Pr(Up|Ctrl) 0.02083333 ## Now in bootstrap.. ## Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred ... ## Point & Interval estimates:
Calculation of risk difference NRI using (‘event’, ‘z.std’, ‘z.std’). nribin(event= event, z.std = z.std, z.new = z.new, cut = c(0.2, 0.4), niter = 1000, updown = 'category') ## STANDARD prediction model: ## Estimate Std. Error z value Pr(>|z|) ## (Intercept) 0.98927136 2.20809035 0.4480212 6.541379e-01 ## age 0.07128234 0.01988079 3.5854876 3.364490e-04 ## bili 0.61686651 0.10992947 5.6114755 2.006087e-08 ## albumin -1.95859156 0.53031693 -3.6932473 2.214085e-04 ## NEW prediction model: ## Estimate Std. Error z value Pr(>|z|) ## (Intercept) -1.16682234 2.92204889 -0.3993165 6.896600e-01 ## age 0.06659224 0.02032242 3.2767864 1.049958e-03 ## bili 0.59995139 0.11022521 5.4429600 5.240243e-08 ## albumin -1.88620553 0.53144647 -3.5491919 3.864153e-04 ## protime 0.20127560 0.18388726 1.0945598 2.737095e-01 ## UP and DOWN calculation: ## #of total, case, and control subjects at t0: 232 88 144 ## Reclassification Table for all subjects: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 110 3 0 ## < 0.4 3 30 0 ## >= 0.4 0 2 84 ## Reclassification Table for case: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 7 0 0 ## < 0.4 0 8 0 ## >= 0.4 0 2 71 ## Reclassification Table for control: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 103 3 0 ## < 0.4 3 22 0 ## >= 0.4 0 0 13 ## NRI estimation: ## Point estimates: ## Estimate ## NRI -0.02272727 ## NRI+ -0.02272727 ## NRI- 0.00000000 ## Pr(Up|Case) 0.00000000 ## Pr(Down|Case) 0.02272727 ## Pr(Down|Ctrl) 0.02083333 ## Pr(Up|Ctrl) 0.02083333 ## Now in bootstrap.. ## Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred ... ## Point & Interval estimates:
Calculation of risk difference NRI using (‘event’, ‘p.std’, ‘p.std’). nribin(event= event, p.std = p.std, p.new = p.new, cut = c(0.2, 0.4), niter = 1000, updown = 'category') ## UP and DOWN calculation: ## #of total, case, and control subjects at t0: 232 88 144 ## Reclassification Table for all subjects: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 110 3 0 ## < 0.4 3 30 0 ## >= 0.4 0 2 84 ## Reclassification Table for case: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 7 0 0 ## < 0.4 0 8 0 ## >= 0.4 0 2 71 ## Reclassification Table for control: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 103 3 0 ## < 0.4 3 22 0 ## >= 0.4 0 0 13 ## NRI estimation: ## Point estimates: ## Estimate ## NRI -0.02272727 ## NRI+ -0.02272727 ## NRI- 0.00000000 ## Pr(Up|Case) 0.00000000 ## Pr(Down|Case) 0.02272727 ## Pr(Down|Ctrl) 0.02083333 ## Pr(Up|Ctrl) 0.02083333 ## Now in bootstrap.. ## Point & Interval estimates:
Calculation of risk difference NRI using (‘mdl.std’, ‘mdl.std’). nribin(mdl.std= mstd, mdl.new = mnew, cut = 0.02, niter = 0, updown = 'diff') ## UP and DOWN calculation: ## #of total, case, and control subjects at t0: 232 88 144 ## #of subjects with 'p.new - p.std > cut' for all, case, control: 34 17 17 ## #of subjects with 'p.std - p.new < cut' for all, case, control: 36 13 23 ## NRI estimation: ## Point estimates:
Calculation of risk difference NRI using (‘event’, ‘z.std’, ‘z.std’). nribin(event= event, z.std = z.std, z.new = z.new, cut = 0.02, niter = 1000, updown = 'diff') ## STANDARD prediction model: ## Estimate Std. Error z value Pr(>|z|) ## (Intercept) 0.98927136 2.20809035 0.4480212 6.541379e-01 ## age 0.07128234 0.01988079 3.5854876 3.364490e-04 ## bili 0.61686651 0.10992947 5.6114755 2.006087e-08 ## albumin -1.95859156 0.53031693 -3.6932473 2.214085e-04 ## NEW prediction model: ## Estimate Std. Error z value Pr(>|z|) ## (Intercept) -1.16682234 2.92204889 -0.3993165 6.896600e-01 ## age 0.06659224 0.02032242 3.2767864 1.049958e-03 ## bili 0.59995139 0.11022521 5.4429600 5.240243e-08 ## albumin -1.88620553 0.53144647 -3.5491919 3.864153e-04 ## protime 0.20127560 0.18388726 1.0945598 2.737095e-01 ## UP and DOWN calculation: ## #of total, case, and control subjects at t0: 232 88 144 ## #of subjects with 'p.new - p.std > cut' for all, case, control: 34 17 17 ## #of subjects with 'p.std - p.new < cut' for all, case, control: 36 13 23 ## NRI estimation: ## Point estimates: ## Estimate ## NRI 0.08712121 ## NRI+ 0.04545455 ## NRI- 0.04166667 ## Pr(Up|Case) 0.19318182 ## Pr(Down|Case) 0.14772727 ## Pr(Down|Ctrl) 0.15972222 ## Pr(Up|Ctrl) 0.11805556 ## Now in bootstrap.. ## Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred ... ## Point & Interval estimates:
Calculation of risk difference NRI using (‘event’, ‘p.std’, ‘p.std’). nribin(event= event, p.std = p.std, p.new = p.new, cut = 0.02, niter = 1000, updown = 'diff') ## UP and DOWN calculation: ## #of total, case, and control subjects at t0: 232 88 144 ## #of subjects with 'p.new - p.std > cut' for all, case, control: 34 17 17 ## #of subjects with 'p.std - p.new < cut' for all, case, control: 36 13 23 ## NRI estimation: ## Point estimates: ## Estimate ## NRI 0.08712121 ## NRI+ 0.04545455 ## NRI- 0.04166667 ## Pr(Up|Case) 0.19318182 ## Pr(Down|Case) 0.14772727 ## Pr(Down|Ctrl) 0.15972222 ## Pr(Up|Ctrl) 0.11805556 ## Now in bootstrap.. ## Point & Interval estimates:
比较新旧两个模型准确性。cut是判断风险高低的临界值,现在我们写了2个,0~20%为低风险,20%~40%为中风险,40%~100%为高风险。根据实际情况确定风险划分的界值,相当于把一个连续的风险概率划分为有序分类变量。updown为定义一个样本的风险是否变动的方式,category是指分类值,即低、中、高风险;另有一种diff,为连续值。选diff时,cut就设1个值,比如0.02,即认为当预测的风险在新旧模型中相差2%时,即被认为是重新分类了。niter为迭代次数,即boostrap重抽样次数,计算NRI的标准误需要重抽样方法计算,如设置为0即表示不计算NRI的标准误,一般迭代次数设置为1000次,如果设置过大对于计算机运算速度要求较高,可能计算要花费很长时间,当然设置越大精确度越高。检验水准alpha一般设为0.05。主要结果截取如下:
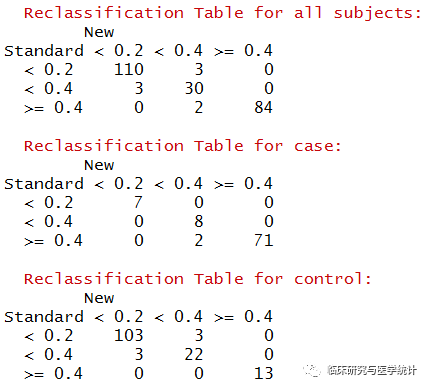
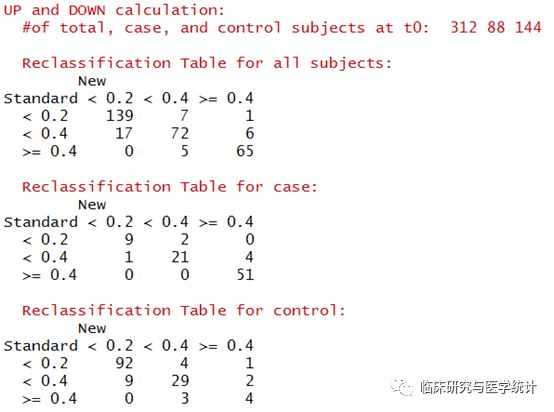
图5. 在所有结局、阳性结局、阴性结局中重分类的表格(case在预测模型中应该理解为结局发生,control理解为结局未发生)
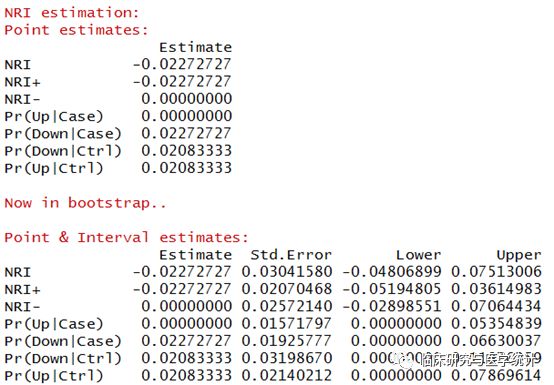
图6. NRI点估计计算结果及重抽样后估计的标准误与可信区间。新模型较旧模型重分类正确的比例提高-2.2727%,换句话说增加一个预测变量的新模型预测的准确度降低了,新模型比旧模型差。 (2)生存资料结局 案例数据同上。生存资料的NRI与分类结局的NRI差别在于前者需要构建Cox回归模型,所以我们首先构建新旧Cox回归模型,计算这两个模型的NRI. R语言代码及结果解读 here consider pbc dataset in survival package as an example library(nricens) dat= pbc[1:312,] dat$sex= ifelse(dat$sex=='f', 1, 0)predciting the event of ‘death’ time= dat$time event= ifelse(dat$status==2, 1, 0)standard prediction model: age, bilirubin, and albumin z.std= as.matrix(subset(dat, select = c(age, bili, albumin)))new prediction model: age, bilirubin, albumin, and protime z.new= as.matrix(subset(dat, select = c(age, bili, albumin, protime)))coxph fit构建Cox生存函数模型 mstd= coxph(Surv(time,event) ~ ., data.frame(time,event,z.std), x=TRUE) mnew= coxph(Surv(time,event) ~ ., data.frame(time,event,z.new), x=TRUE)predicted risk at t0=2000,2000天时间点的死亡风险 p.std= get.risk.coxph(mstd, t0=2000) p.new= get.risk.coxph(mnew, t0=2000)以上新、旧Cox回归模型构建完成,并计算了2000天的新、旧模型计算的死亡风险 Calculation of risk category NRI by the KM estimator using (‘mdl.std’, ‘mdl.std’). nricens(mdl.std= mstd, mdl.new = mnew, t0 = 2000, cut = c(0.2, 0.4), niter = 1000, updown = 'category') ## UP and DOWN calculation: ## #of total, case, and control subjects at t0: 312 88 144 ## Reclassification Table for all subjects: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 139 7 1 ## < 0.4 17 72 6 ## >= 0.4 0 5 65 ## ## Reclassification Table for case: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 9 2 0 ## < 0.4 1 21 4 ## >= 0.4 0 0 51 ## Reclassification Table for control: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 92 4 1 ## < 0.4 9 29 2 ## >= 0.4 0 3 4 ## NRI estimation by KM estimator: ## Point estimates: ## Estimate ## NRI 0.11028068 ## NRI+ 0.05123381 ## NRI- 0.05904686 ## Pr(Up|Case) 0.06348538 ## Pr(Down|Case) 0.01225156 ## Pr(Down|Ctrl) 0.09583016 ## Pr(Up|Ctrl) 0.03678329 ## Now in bootstrap.. ## Point & Interval estimates:
by the KM estimator using (‘time’, ‘event’, ‘z.std’, ‘z.std’). nricens(time= time, event = event, z.std = z.std, z.new = z.new, t0 = 2000, cut = c(0.2, 0.4), niter = 1000,updown = 'category') ## STANDARD prediction model (Cox model): ## coef exp(coef) se(coef) z Pr(>|z|) ## age 0.03726683 1.0379699 0.009048925 4.118371 3.815600e-05 ## bili 0.13531179 1.1448937 0.013711323 9.868617 5.694436e-23 ## albumin -1.44611854 0.2354825 0.221997986 -6.514107 7.312356e-11 ## NEW prediction model (Cox model): ## coef exp(coef) se(coef) z Pr(>|z|) ## age 0.03362675 1.0341985 0.009214173 3.649460 2.627925e-04 ## bili 0.12517886 1.1333511 0.014406820 8.688861 3.660902e-18 ## albumin -1.39395237 0.2480928 0.217046959 -6.422354 1.341831e-10 ## protime 0.28602917 1.3311313 0.070536400 4.055058 5.012193e-05 ## UP and DOWN calculation: ## #of total, case, and control subjects at t0: 312 88 144 ## Reclassification Table for all subjects: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 139 7 1 ## < 0.4 17 72 6 ## >= 0.4 0 5 65 ## Reclassification Table for case: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 9 2 0 ## < 0.4 1 21 4 ## >= 0.4 0 0 51 ## Reclassification Table for control: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 92 4 1 ## < 0.4 9 29 2 ## >= 0.4 0 3 4 ## ## NRI estimation by KM estimator: ## Point estimates: ## Estimate ## NRI 0.11028068 ## NRI+ 0.05123381 ## NRI- 0.05904686 ## Pr(Up|Case) 0.06348538 ## Pr(Down|Case) 0.01225156 ## Pr(Down|Ctrl) 0.09583016 ## Pr(Up|Ctrl) 0.03678329 ## Now in bootstrap.. ## Point & Interval estimates:
by the KM estimator using (‘time’,‘event’,‘p.std’,‘p.std’). nricens(time= time, event = event, p.std = p.std, p.new = p.new, t0 = 2000, cut = c(0.2, 0.4), niter = 1000,updown = 'category') ## UP and DOWN calculation: ## #of total, case, and control subjects at t0: 312 88 144 ## Reclassification Table for all subjects: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 139 7 1 ## < 0.4 17 72 6 ## >= 0.4 0 5 65 ## Reclassification Table for case: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 9 2 0 ## < 0.4 1 21 4 ## >= 0.4 0 0 51 ## Reclassification Table for control: ## New ## Standard < 0.2 < 0.4 >= 0.4 ## < 0.2 92 4 1 ## < 0.4 9 29 2 ## >= 0.4 0 3 4 ## NRI estimation by KM estimator: ## Point estimates: ## Estimate ## NRI 0.11028068 ## NRI+ 0.05123381 ## NRI- 0.05904686 ## Pr(Up|Case) 0.06348538 ## Pr(Down|Case) 0.01225156 ## Pr(Down|Ctrl) 0.09583016 ## Pr(Up|Ctrl) 0.03678329 ## Now in bootstrap.. ## Point & Interval estimates:
Calculation of risk difference NRI by the KM estimator nricens(mdl.std= mstd, mdl.new = mnew, t0 = 2000, updown = 'diff', cut = 0.05, niter = 1000) ### UP and DOWN calculation: ## #of total, case, and control subjects at t0: 312 88 144 ## #of subjects with 'p.new - p.std > cut' for all, case, control: 34 21 11 ## #of subjects with 'p.std - p.new < cut' for all, case, control: 40 12 8 ## NRI estimation by KM estimator: ## Point estimates: ## Estimate ## NRI 0.10070960 ## NRI+ 0.05097223 ## NRI- 0.04973737 ## Pr(Up|Case) 0.22431499 ## Pr(Down|Case) 0.17334277 ## Pr(Down|Ctrl) 0.10859064 ## Pr(Up|Ctrl) 0.05885327 ## Now in bootstrap.. ## Point & Interval estimates:
Calculation of risk difference NRI by the IPW estimator nricens(mdl.std= mstd, mdl.new = mnew, t0 = 2000, updown = 'diff', cut = 0.05, pointhod = 'ipw', niter= 1000) ## UP and DOWN calculation: ## #of total, case, and control subjects at t0: 312 88 144 ## #of subjects with 'p.new - p.std > cut' for all, case, control: 34 21 11 ## #of subjects with 'p.std - p.new < cut' for all, case, control: 40 12 8 ## NRI estimation by IPW estimator: ## Point estimates: ## Estimate ## NRI 0.06361038 ## NRI+ 0.08444371 ## NRI- -0.02083333 ## Pr(Up|Case) 0.22905909 ## Pr(Down|Case) 0.14461537 ## Pr(Down|Ctrl) 0.05555556 ## Pr(Up|Ctrl) 0.07638889 ## Now in bootstrap.. ## Point & Interval estimates:
参数解释同分类结局。 主要计算结果如下图7.与图8.所示:
图7. 在所有结局、阳性结局、阴性结局中重分类的表格(case在Cox预测模型中应该理解为结局发生,control理解为结局未发生)
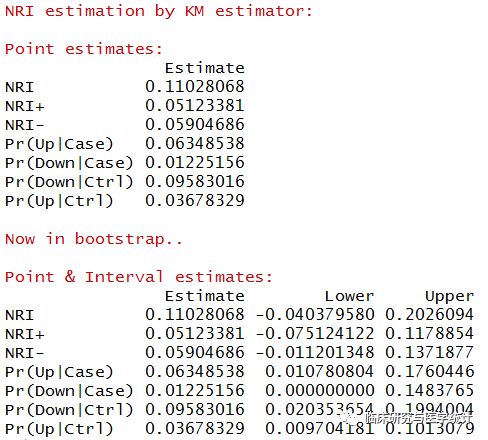
图8. NRI点估计的计算结果及重抽样后估计的可信区间。新模型较旧模型重分类正确的比例提高11.028%,换句话说增加一个预测变量的新模型预测的准确度比旧模型更好。 本章关于NRI的计算讲解完毕。下一章我们将介绍另外一个指标IDI(Integrated Discrimination Improvement,综合判别改善指数)的计算原理与计算方法,欢迎关注。 4. 参考文献 [1] Alba A C, Agoritsas T, Walsh M, et al.Discrimination and Calibration of Clinical Prediction Models: Users’ Guides tothe Medical Literature [J]. Jama, 2017, 318(14): 1377-84. [2] Pencina M J, D’Agostino R B, Sr., D’AgostinoR B, Jr., et al. Evaluating the added predictive ability of a new marker: fromarea under the ROC curve to reclassification and beyond [J]. Statistics inmedicine, 2008, 27(2): 157-72; discussion 207-12. [3] http://ncook.bwh.harvard.edu/r-code.html [4] Pencina MJ, D’Agostino RB, Steyerberg EW. Extensions of net reclassificationimprovement calculations to measure usefulness of new biomarkers. Statistics inMedicine 2011. [5] Uno H, Tian L, Cai T, Kohane IS, Wei LJ. A unified inferenceprocedure for a class of measures to assess improvement in risk predictionsystems with survival data, Statistics in Medicine 2012. [6] Hsu CH, Taylor JMG. A robust weighted Kaplan-Meier approach fordata with dependent censoring using linear combinations of prognosticcovariates, Statistics in Medicine 2010. |
【本文地址】
今日新闻 |
推荐新闻 |